Our mission is to enhance biodiversity and restore ecosystems through the genetic rescue of endangered and extinct species. Revive & Restore is the leading wildlife conservation organization promoting the incorporation of biotechnologies into standard conservation practice.
BRINGING BIOTECHNOLOGIES TO CONSERVATION
Latest News
Featured projects
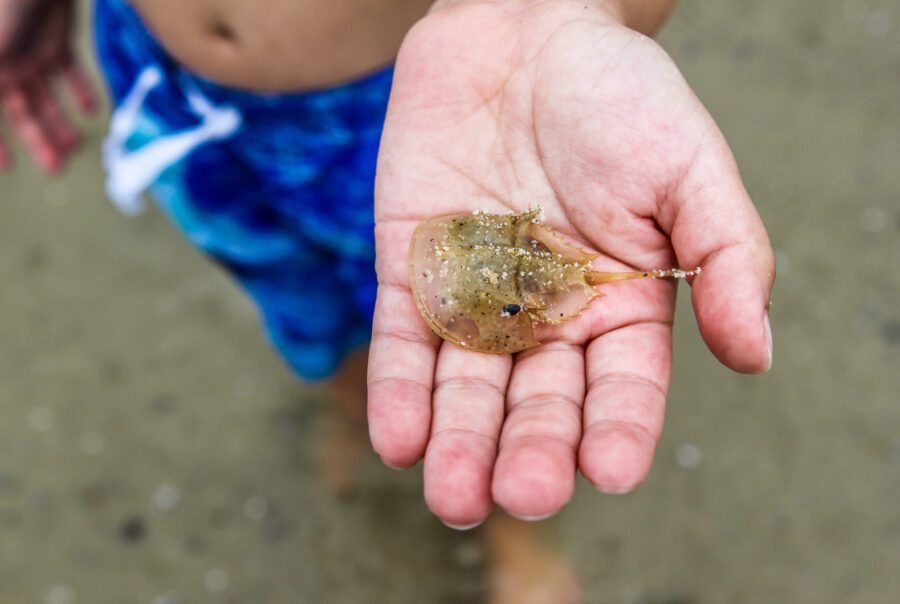

Advancing alternatives to horseshoe crab blood
Every year, hundreds of thousands of horseshoe crabs are harvested for their blood, despite the availability of safe and effective alternatives. We advocate for the adoption of synthetic alternatives to blue blood for endotoxin testing.

Our 2023 Annual Report:
Revive & Restore On the move
This year, we pushed the boundaries of conservation and spearheaded groundbreaking research. Our 2023 Annual Report offers an overview of the milestones we’ve achieved together and the impact your support can have on wildlife.
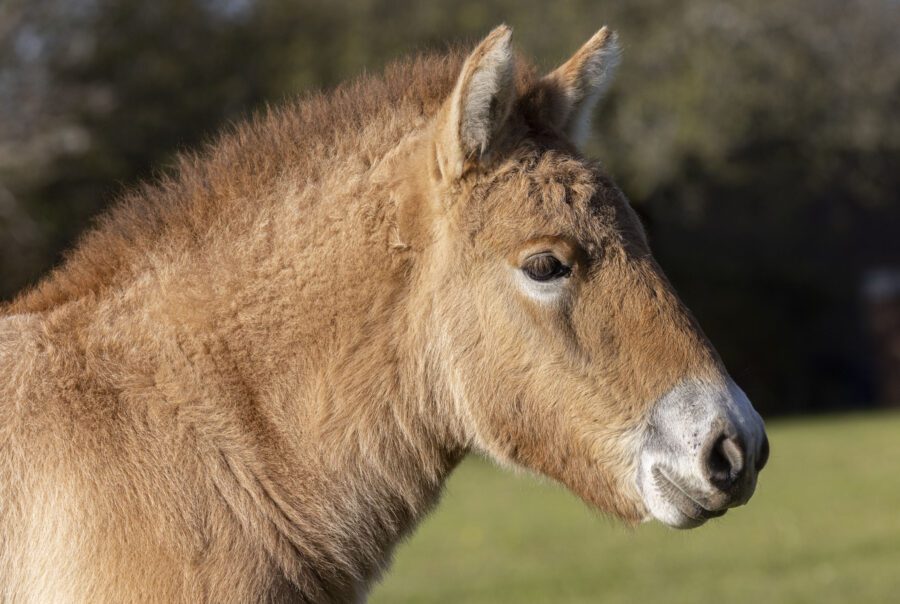

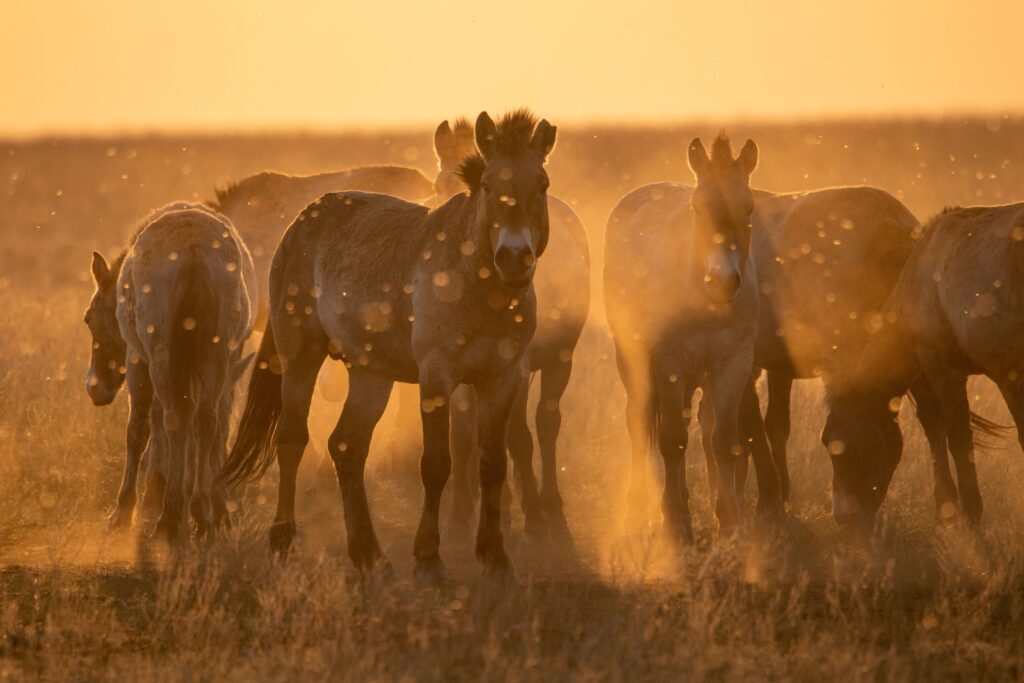
Meet Ollie, our second Przewalski’s horse clone
We’re excited to announce the birth of the world’s second successfully cloned Przewalski’s horse! The new foal, born February 17th, offers hope that cloning can be a viable tool for the genetic rescue of endangered species.
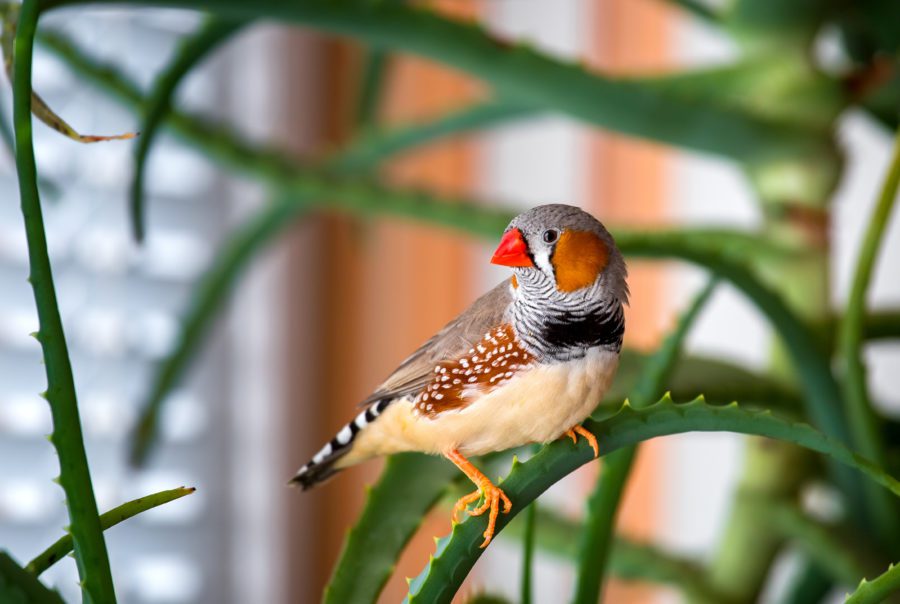
BIOTECHnology FOR BIRD CONSERVATION
The newly launched “Biotechnology for Bird Conservation Program” will enable genetic rescue biotechnologies to help save endangered birds. Awarded projects from our first call for proposals have been announced.
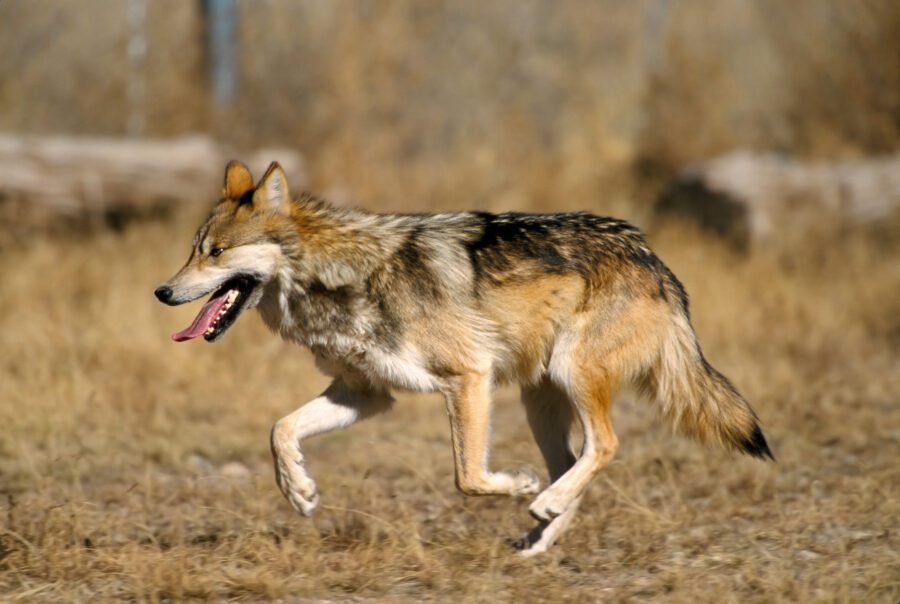

INFORMED Biobanking FOR U.S. endangered species
Revive & Restore, in partnership with the U.S. Fish and Wildlife Service, has embarked on a bold endeavor to biobank U.S. endangered species. This is the first time the USFWS has partnered on an agency-wide biobanking initiative.
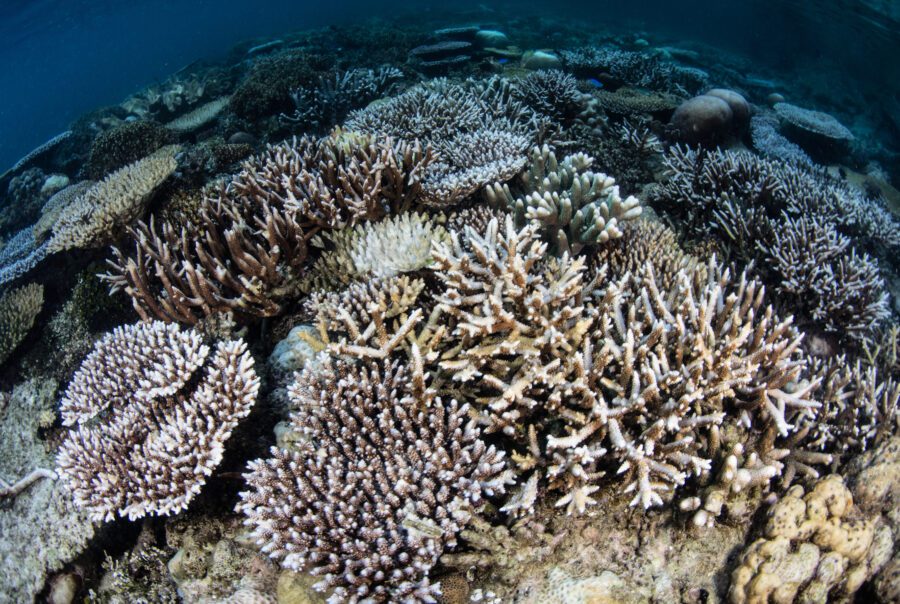

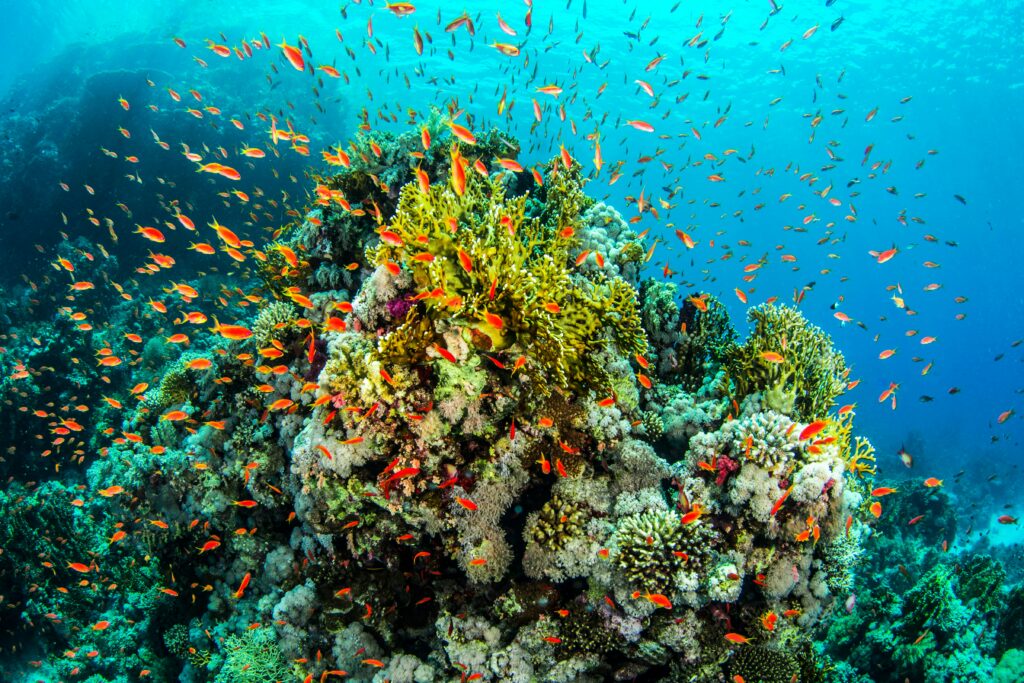
Discover the Advanced Coral Toolkit
The Advanced Coral Toolkit is a funding program that supports the development and field testing of new biotechnologies that have the potential to greatly benefit coral resilience and restoration efforts.
Why this work matters
Together, we can reverse the extinction trajectory.
Biotechnology offers tools that can enhance conservation outcomes, from restoring genetic diversity to facilitating adaptation. Together, we CAN turn the tide on species loss.